Tally-2.3 - Documentation
Tally-2.3 is a scoring tool based on a machine learning approach, which allows to validate the results of tandem repeat detection in protein sequences.
- 1. Input format
- 1.1 MSA only
- 1.2 FASTA
- 1.3 Fastally
- 2. Score
- Tally
- Psim
- p-value-phylo
- Entropy
- Parsimony
- 3. Example
- 3.1 Input
- 3.2 Output
1. Input format
1.1 MSA only
Insert MSA of repeats from one tandem repeat region like below :
Example :
1.2 FASTA
Insert MSA of repeats from one tandem repeat region in FASTA format.
Example :
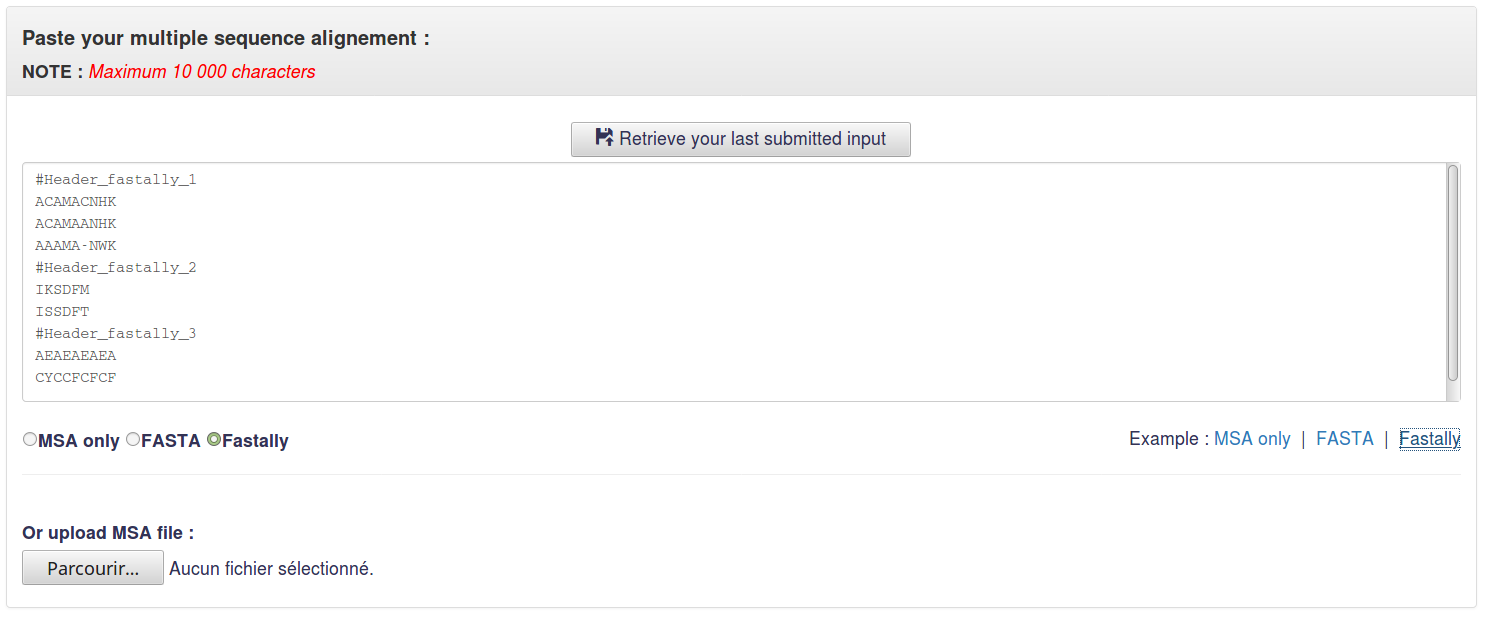
1.3 Fastally
Fastally format allows user to analyse several MSAs at one run.
In this format, headers started with # symbol separate MSAs of different tandem repeat regions.
Example :
2. Score
Tally
Tally-2.3 score is obtained with machine learning approach. At a threshold of 0.5, established based on the maximization of F-score,
Tally-2.3 performs at a level of 89% sensitivity, while achieving a high specificity of 89% and an Area Under the Receiver Operating Characteristic Curve of 96%.
Validated MSAs have Tally-2.3 scores ≥ 0.5.
Psim
Psim is a score relying on the Hamming distance between the repeats and their consensus sequence.
Validated MSAs have Psim ≥ 0.7. [Psim documentation]
p-value-phylo
Validated MSAs have p-value-phylo scores ≤ 0.001. [Schaper et al.,2012]
Entropy
See : Entropy score definition
Parsimony
See : Parsimony score definition
3. Example
3.1 Input
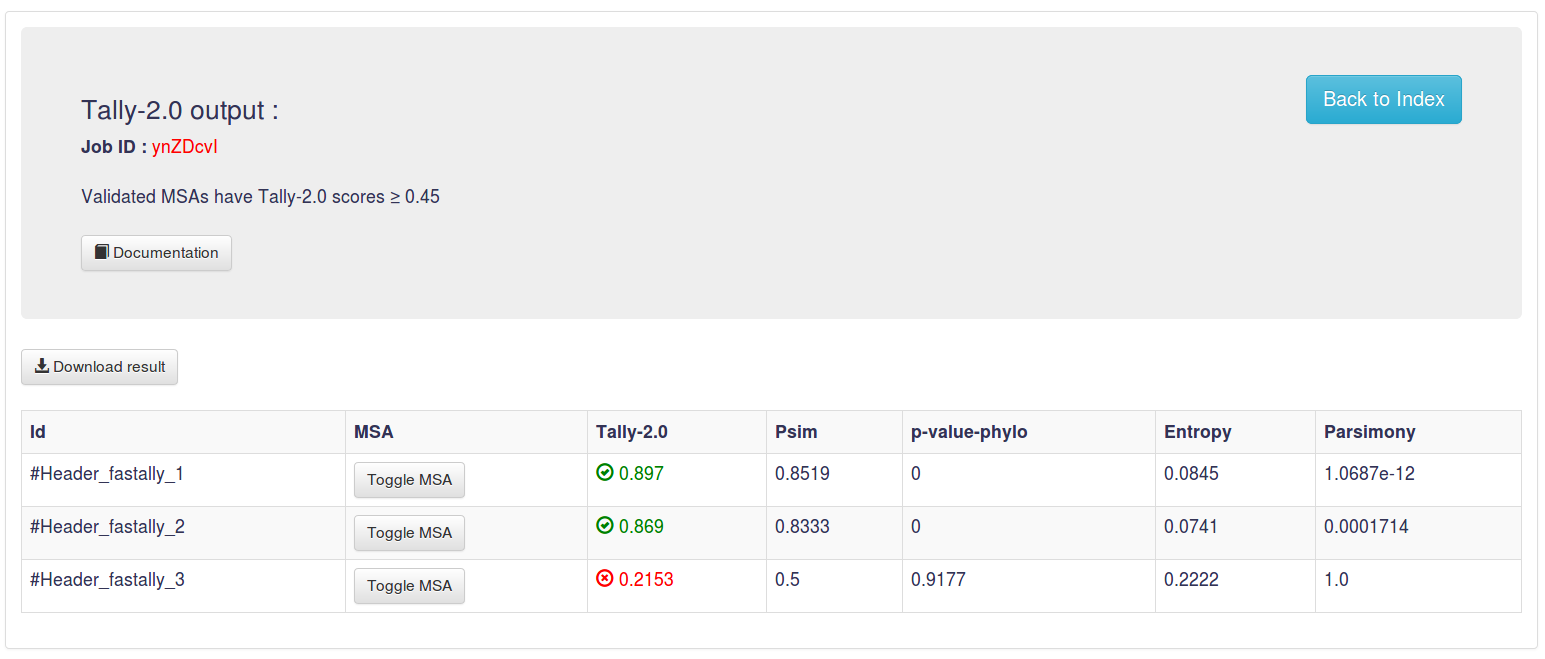
3.2 Output